Cancer research and pathology relies heavily on formalin-fixed, paraffin embedded (FFPE) tissues from patients. While FFPE samples are easy and cost-effective to store, they are often incompatible with contemporary omics techniques, like single-cell sequencing.
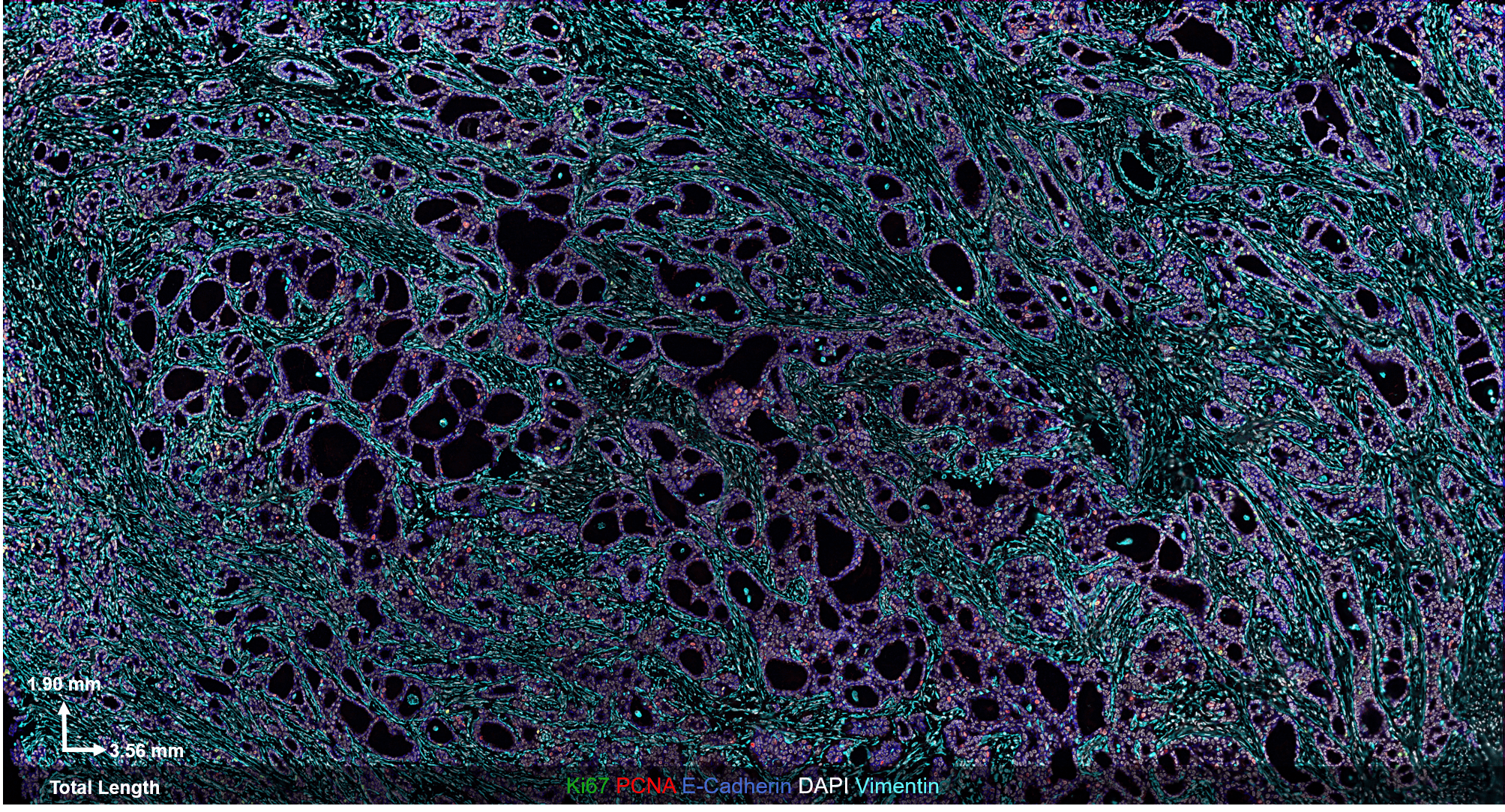
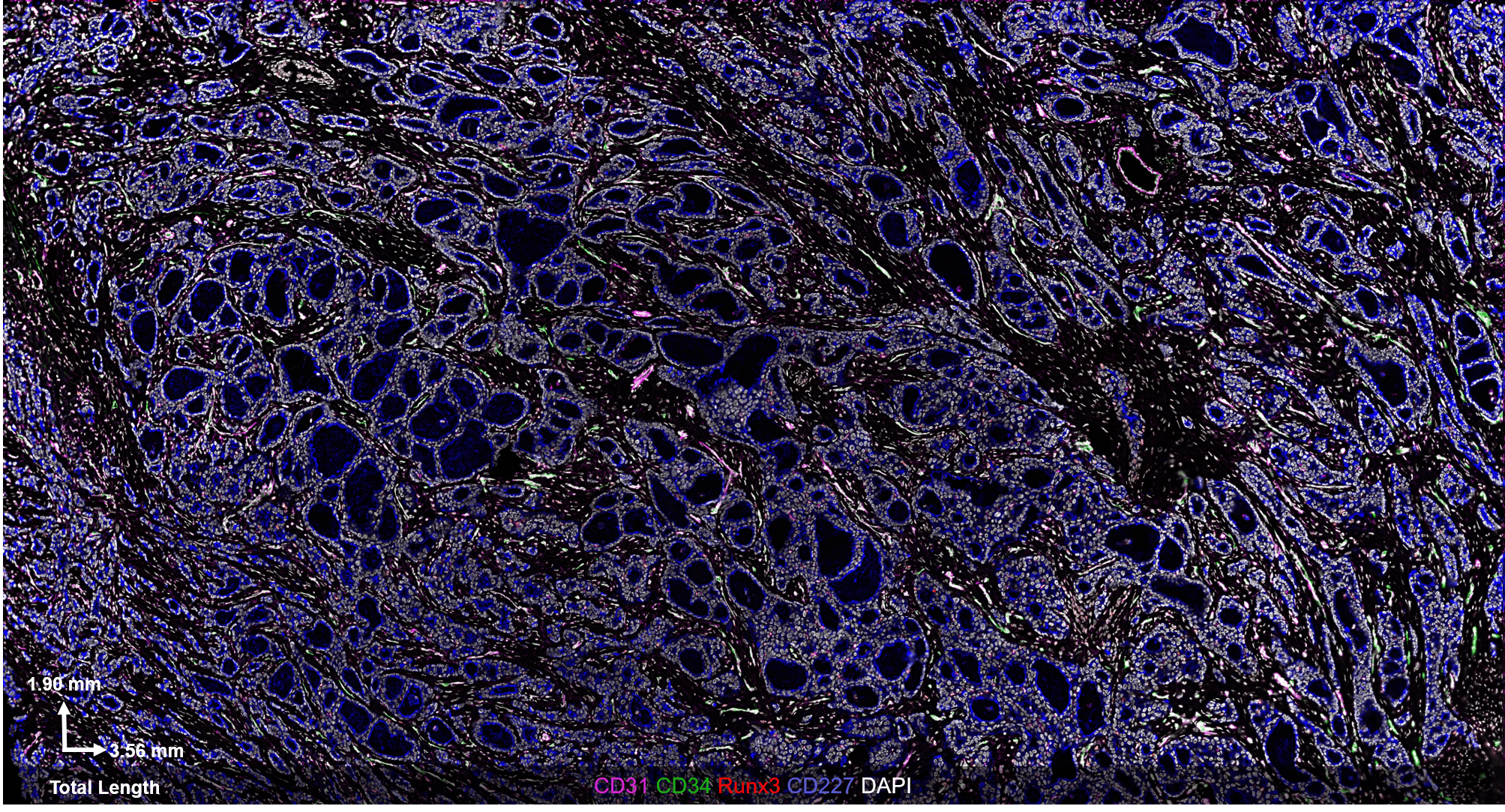
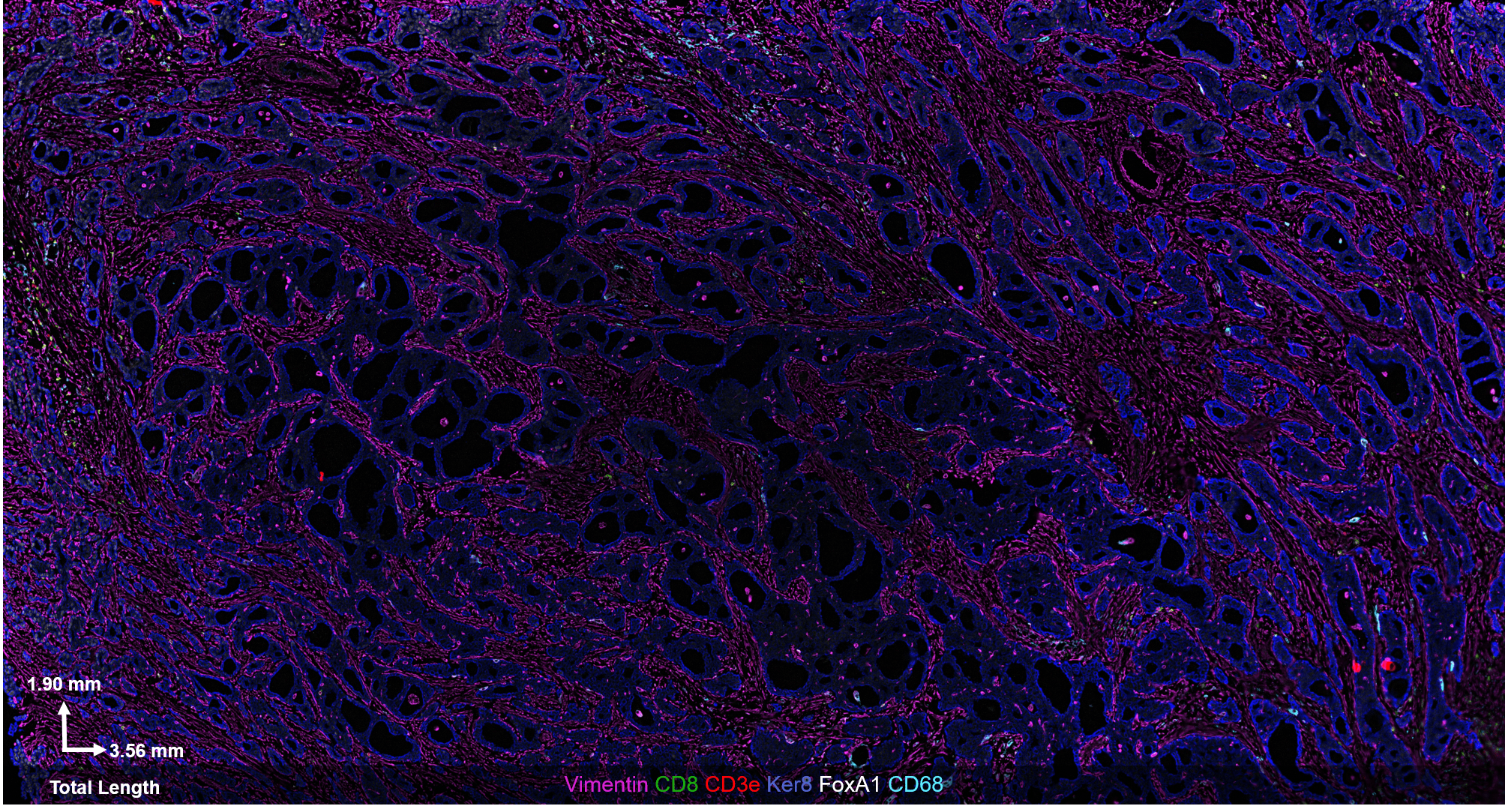
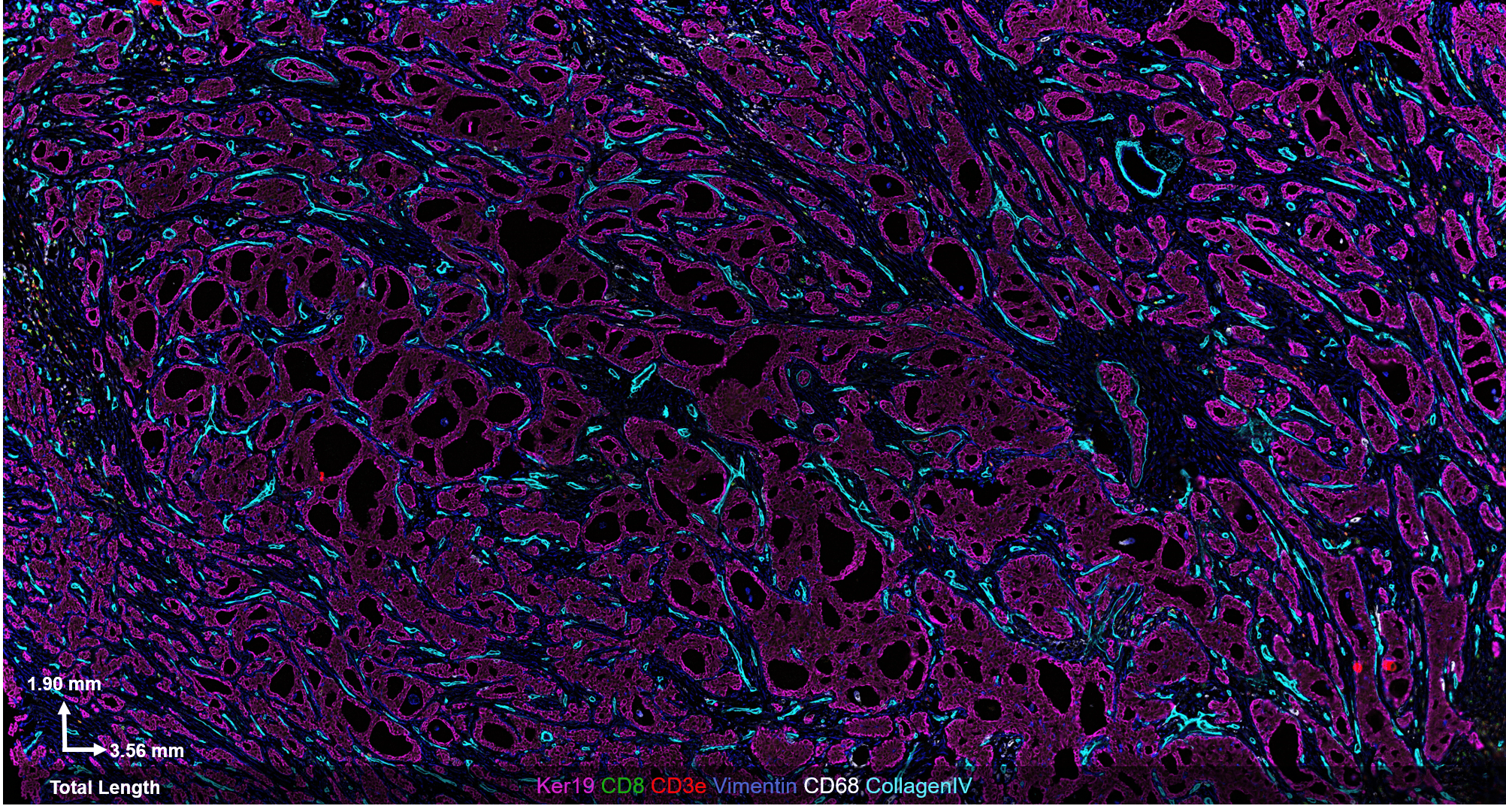
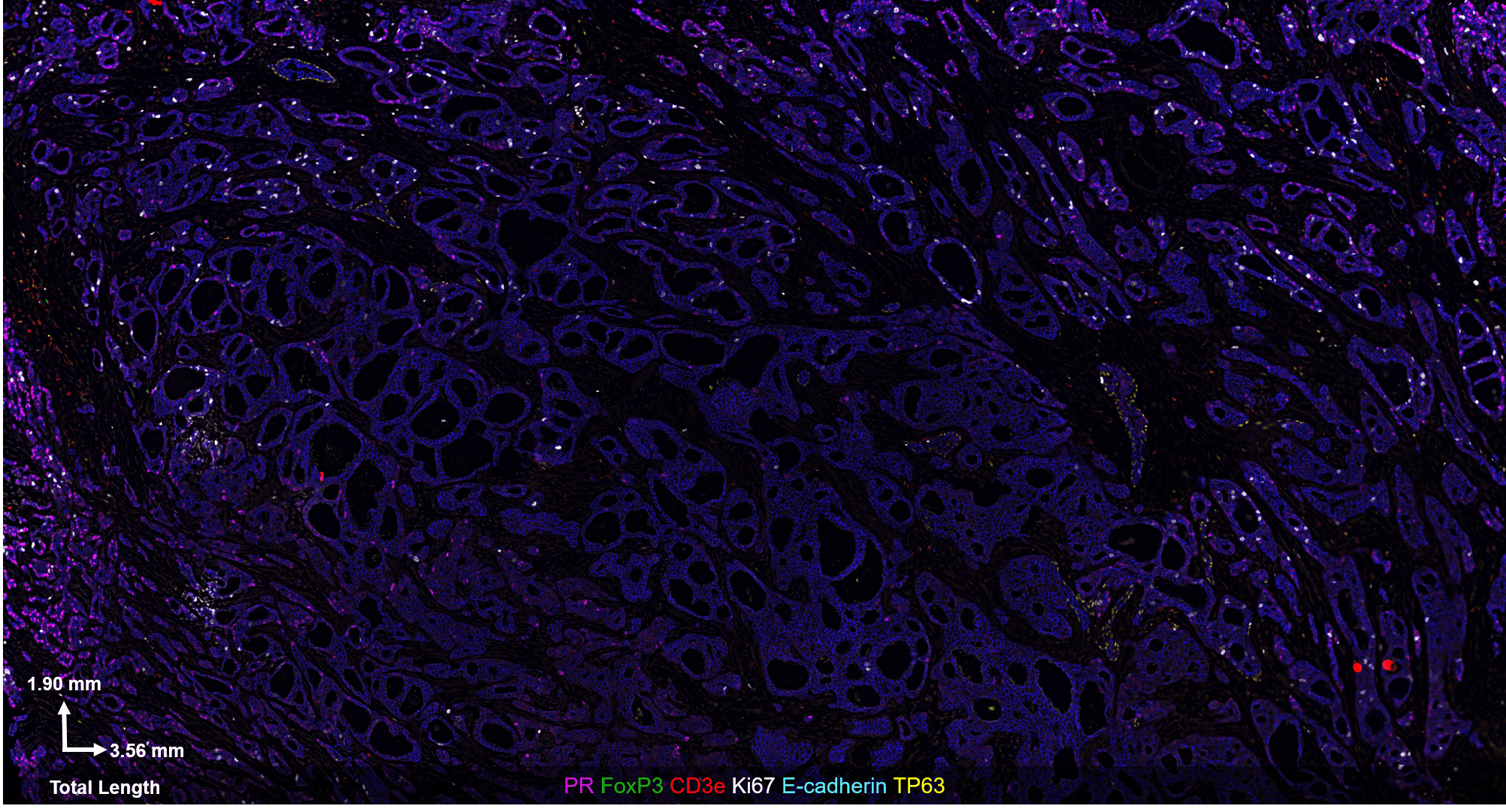
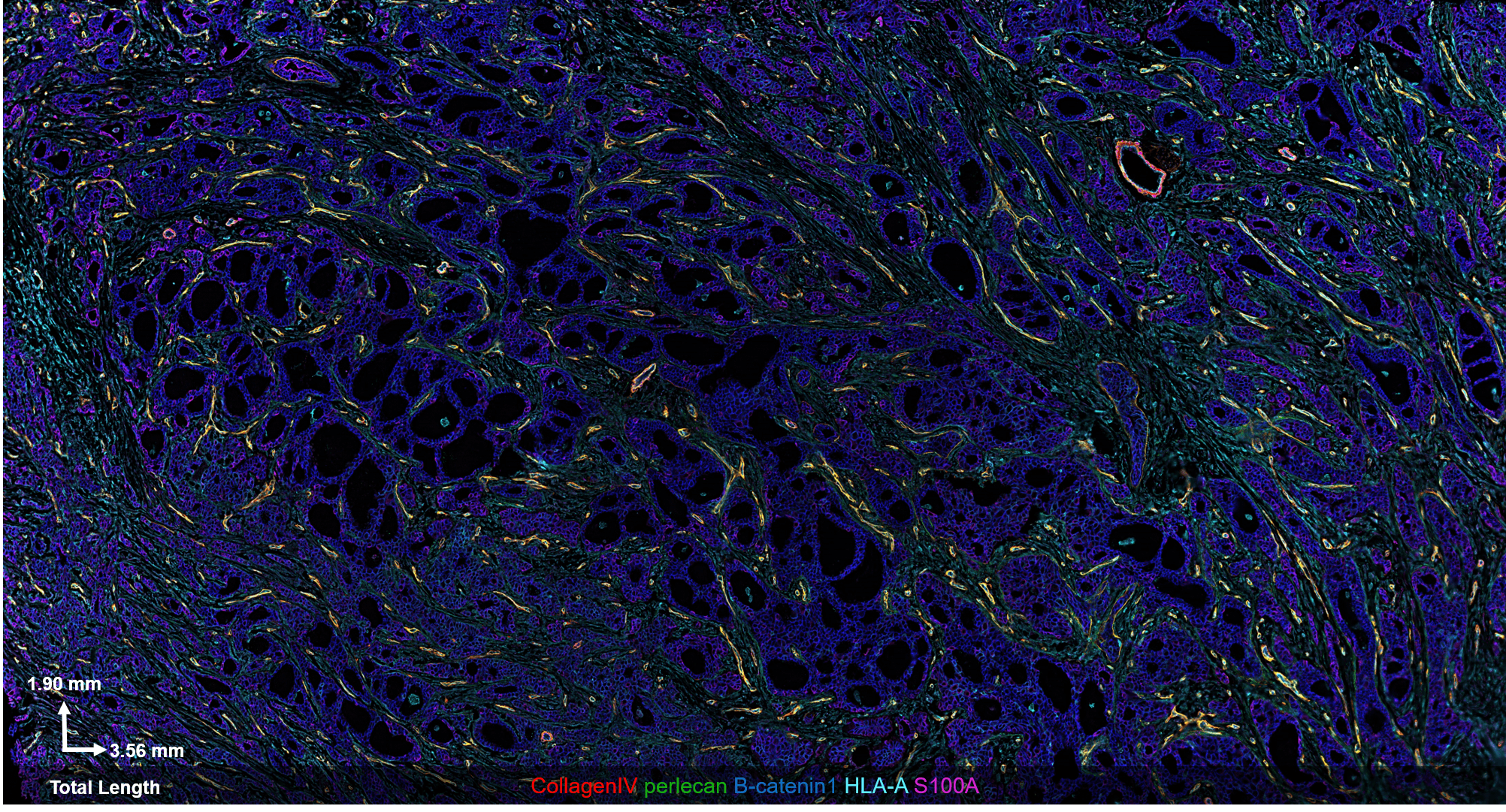
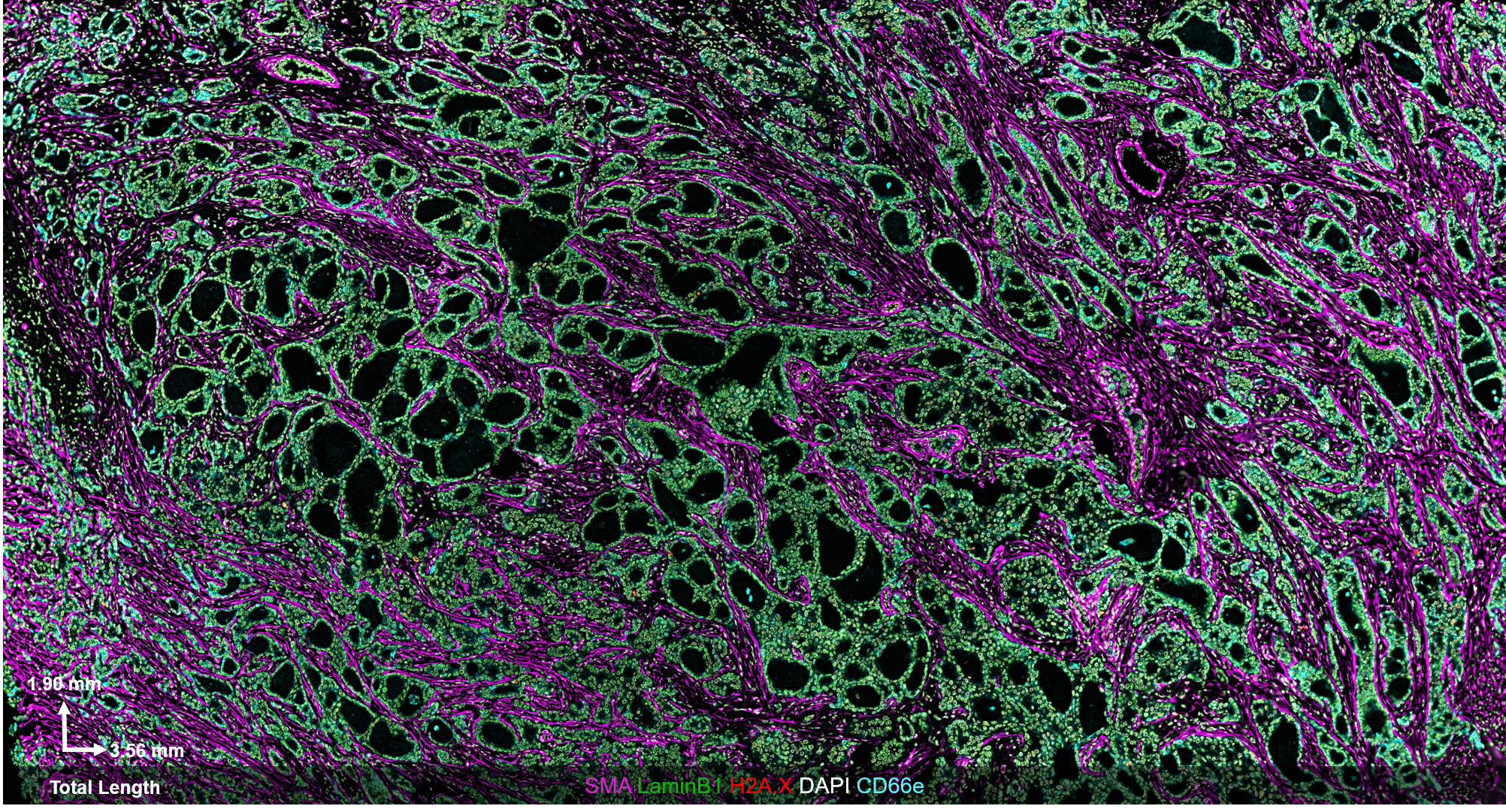
The CODEX® platform overcomes this limitation. It’s compatible with FFPE tissues and can image 40+ protein biomarkers with single-cell resolution, while retaining spatial context. The Akoya team developed and tested the following antibody panel to study breast cancer in human FFPE tissue using CODEX. This ultra-high plex, 36-antibody panel contains mostly epithelial markers targeted at different secretory and non-secretory epithelia in breast tissue, as well as cell state and proliferation markers that are frequently implicated in cancer etiology.
Panel Information
Tissue: Stage II Ductal Adenocarcinoma
Plex: 36-Antibodies with many of them designed to detect epithelial components
Size: 3.56mm x 1.90mm or 6.76mm2
List of Markers:
CD31
CD227
Collagen IV
E-cadherin
H2A.X
Ker 5
Ker 8
Ker 14
Ker 17
Ker18
Ker 19
LaminB1
Myosin
perlecan
S100A4
SMA
Vimentin
B-catenin1
CD66e
E2F1
FoxA1
p53
TFAM
CD3e
CD8
CD68
FOXP3
B-actin
CD34
Ki67
MHCI
MHCII
PCNA
PR
RUNX3
Tp63
Recapitulation of independently published findings
We compared our results to a Nature Communications study to demonstrate their reliability. In the paper, “Profiling human breast epithelial cells using single cell RNA sequencing identifies cell diversity”, Nguyen et. al. use single-cell RNA sequencing and low-plex histology to produce an atlas of ductal epithelial cells in normal breast tissue samples. They describe two main cell types: basal and luminal epithelia.
Our team independently replicated two of their observations in FFPE Stage II Ductal Adenocarcinoma.
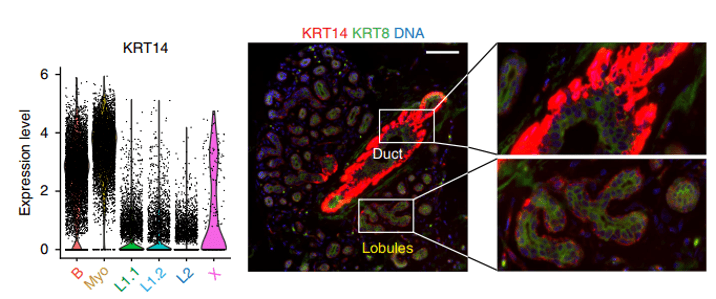
Fig 1. Results from Nature Communications paper. Image Source: Nguyen et al (2018) / CC-BY-NC-ND 4.0.
The figure above shows a violin plot (left) and imaging data (right) from the normal breast epithelium. The epithelium can be separated into ducts and lobules, which serve different functions. Nguyen et. al. show that KRT14 is strongly expressed in Basal and Myoepithelial cells, which are shown in red in the imaging data. Lower KRT14 expression is observed in luminal L1 and L2 cells. These are shown in green, cyan and blue.
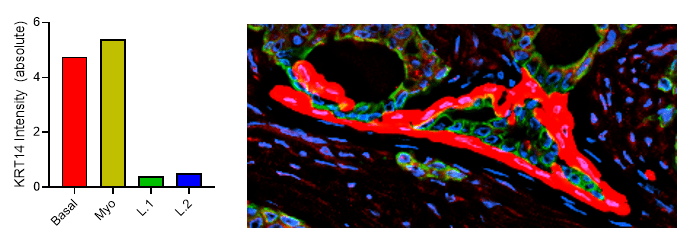
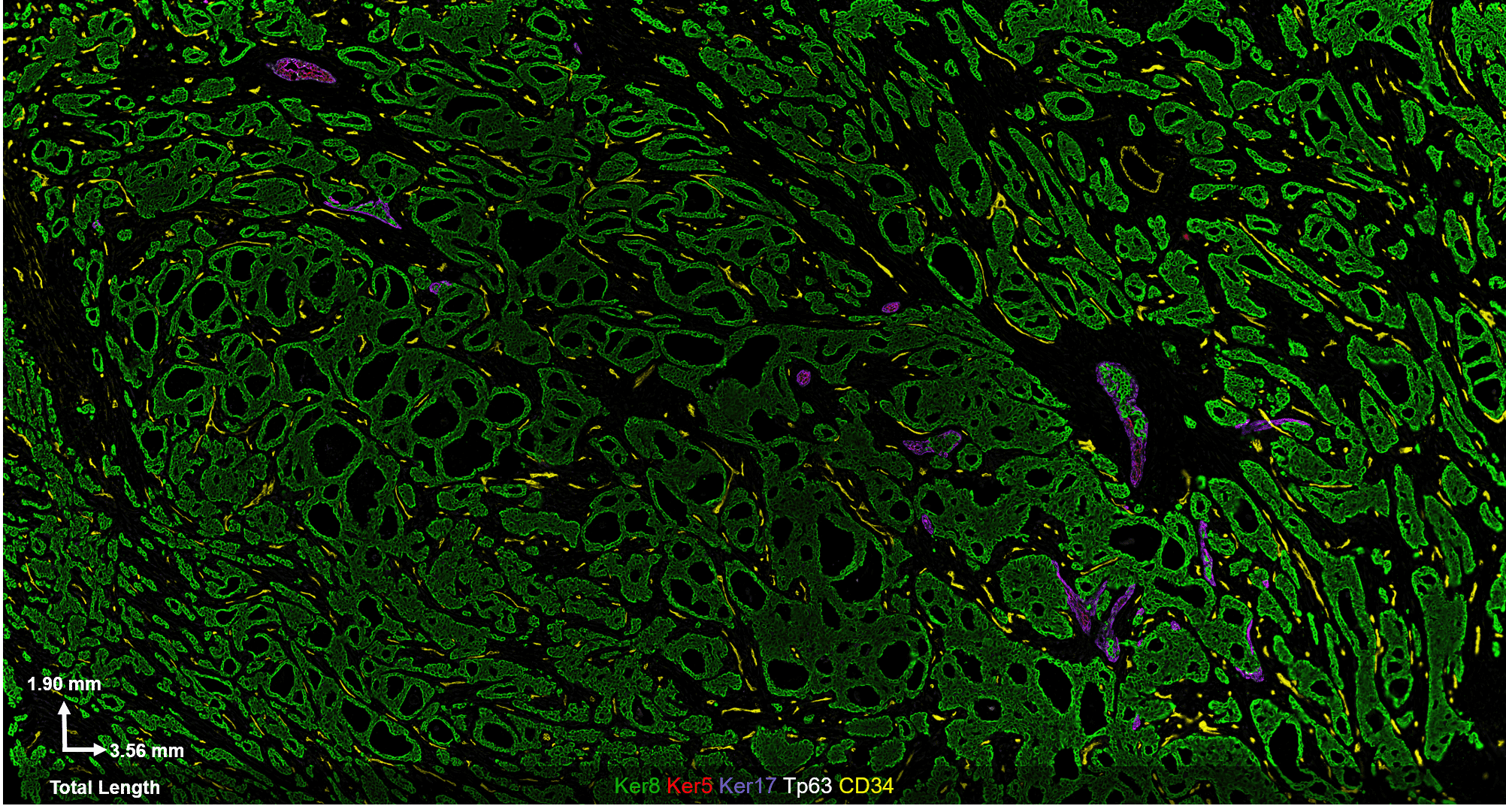
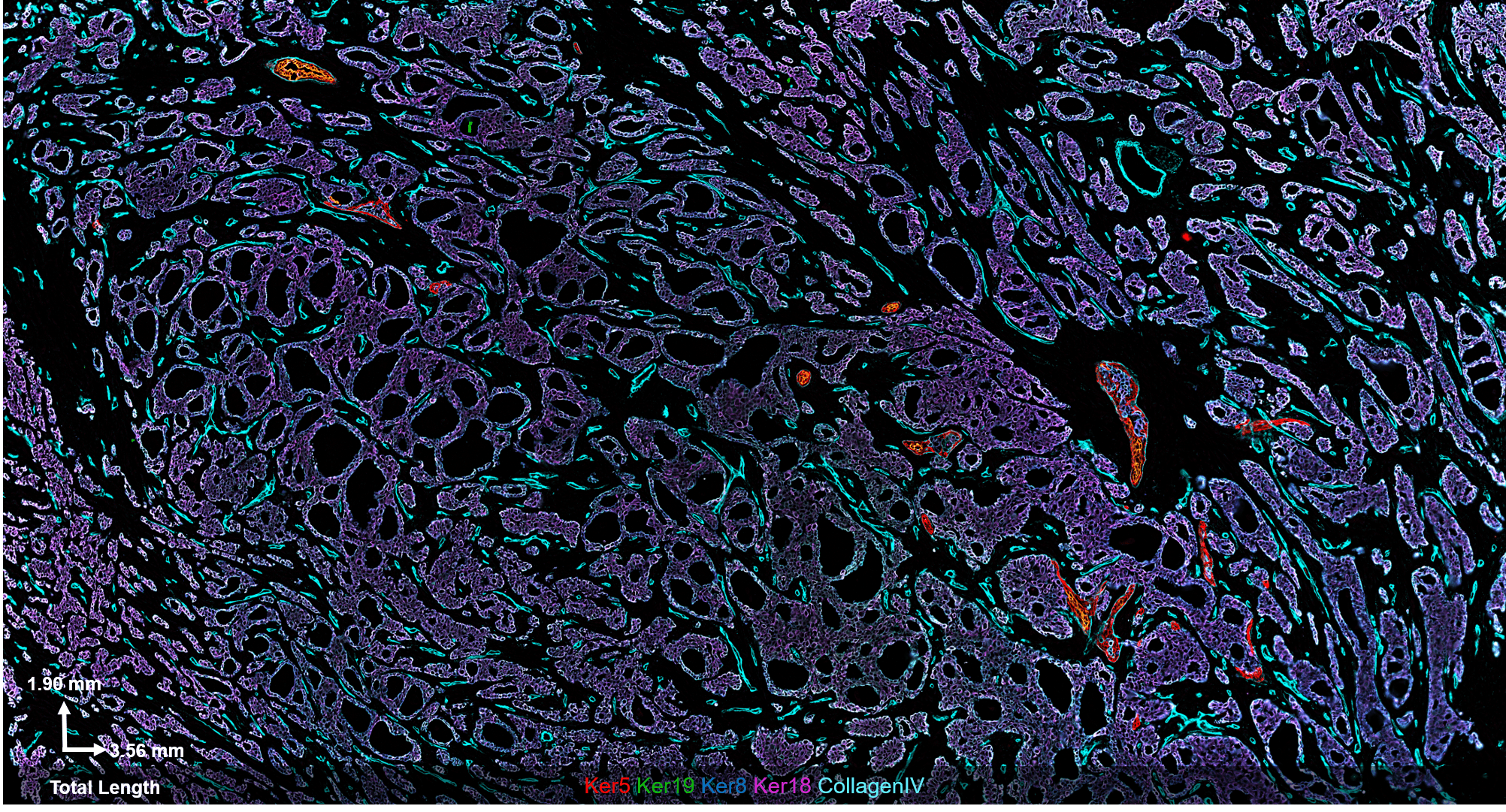
Fig 2. Results from Akoya’s Internal Analysis
Our results were obtained with a CODEX 36 antibody breast panel on Human Stage II Ductal Adenocarcinoma, and they showed excellent recapitulation of the published results. We see the KRT14 (red) being strongly expressed in the Ductal Basal and Myo-epithelium. In the lobules – luminal cells, L.1 and L.2 in the histogram – there is much lower KRT14 expression.
This result was replicated without intent and in a different tissue sample/patient condition, which speaks to the reliability and robustness of our experiment. CODEX produces reliable staining over a range of biomarker expression intensities.
How to get started
Interested in learning more about this panel? Check out this on-demand webinar where our Senior Applications Scientist, Dr. Oliver Braubach, covers the results of this experiment in more depth.
We’ve also compiled a helpful ordering list of the CODEX Antibodies and CODEX Barcode products that can be used to analyze biomarkers shown in this panel. Contact your local sales rep to purchase the reagents. If you’re new to CODEX, contact us to get started.